New Publication: Exploring the proteome and phosphoproteome response to mistranslation
Our latest paper was recently published in G3: Genes, Genomes, Genetics. You can check out the publication here.
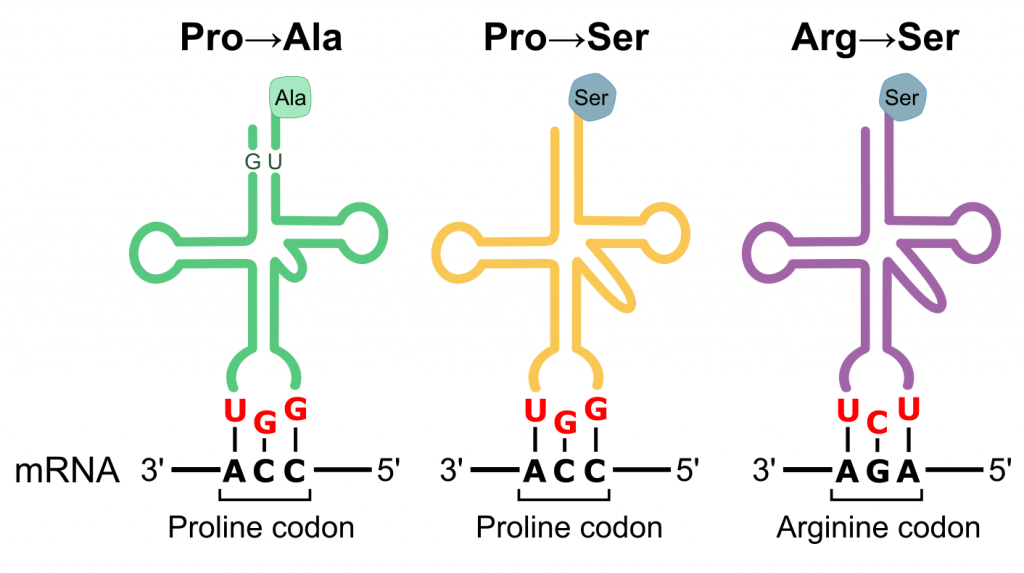
In this paper, we used quantitative proteomics and phosphoproteomics to profile the cellular response of yeast to three different mistranslating tRNA variants.
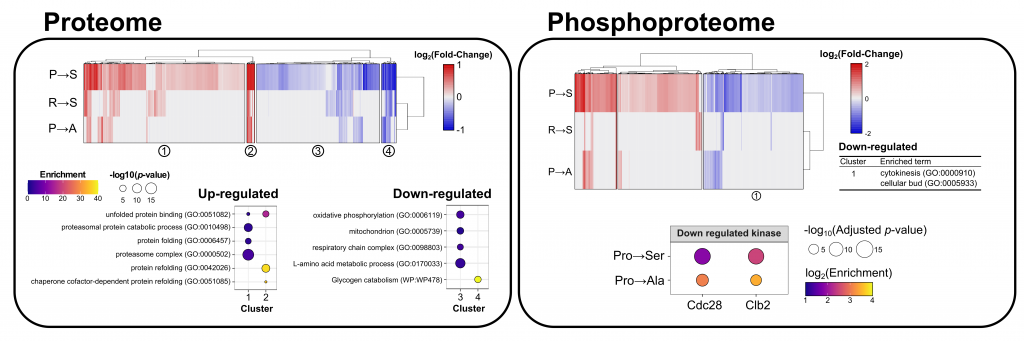
We find that although global proteome impacts were modest, mistranslating tRNAs affect protein abundance and phosphorylation in key pathways including proteostasis, cell cycle regulation and translation.
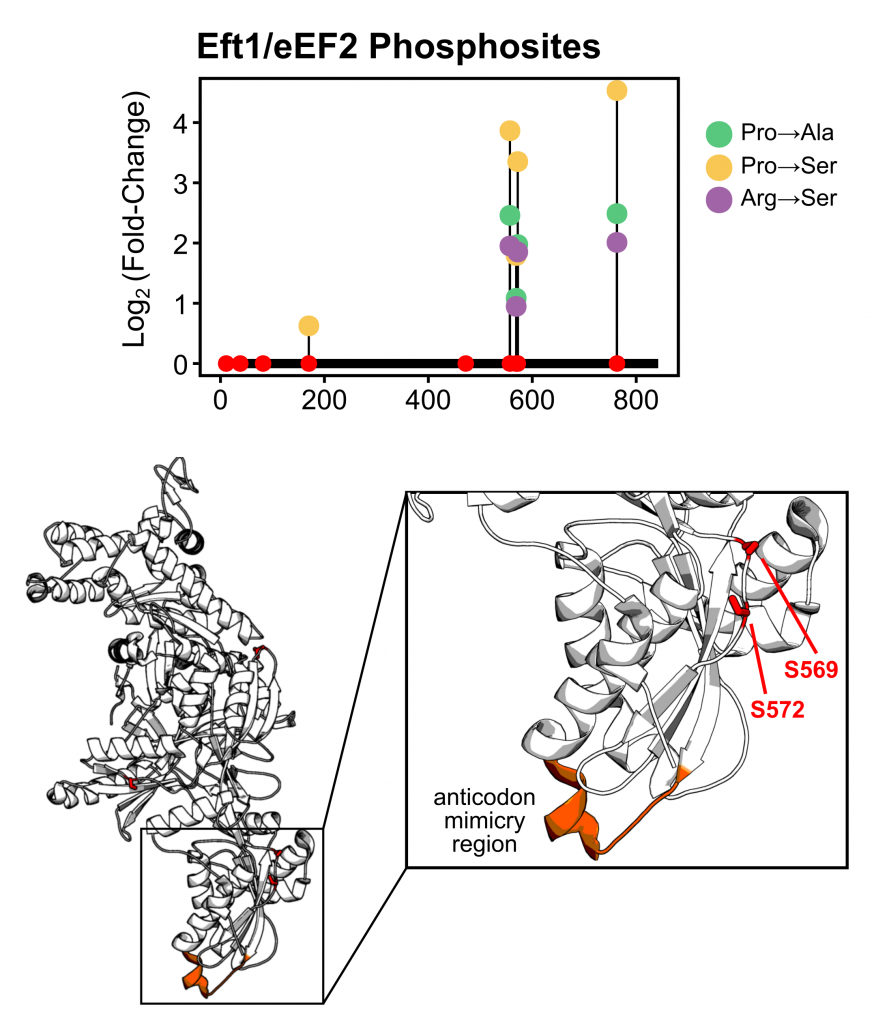
Our dataset uncovers many changes, particularly in the phosphoproteome, that merit further investigation and provides a resource for understanding how mistranslation shapes cellular pathways. One particularly exciting finding is the upregulation of phosphosites on proteins involved in translation. Specifically, the most upregulated phosphosites occurred on eEF2 which could potentially suggest a link between mistranslation and translation speed. It will be exciting to follow up on these sites to determine their functional impact.
We also find new phosphosites created by mistranslating serine at non-serine positions and we hypothesize that they could impact protein function.
Overall, these findings underscore the importance of considering cellular consequences when engineering tRNAs for therapeutic applications and will be of interest to researchers studying tRNA biology, proteostasis and regulation of translation fidelity.